Light and shade of TCR promiscuity
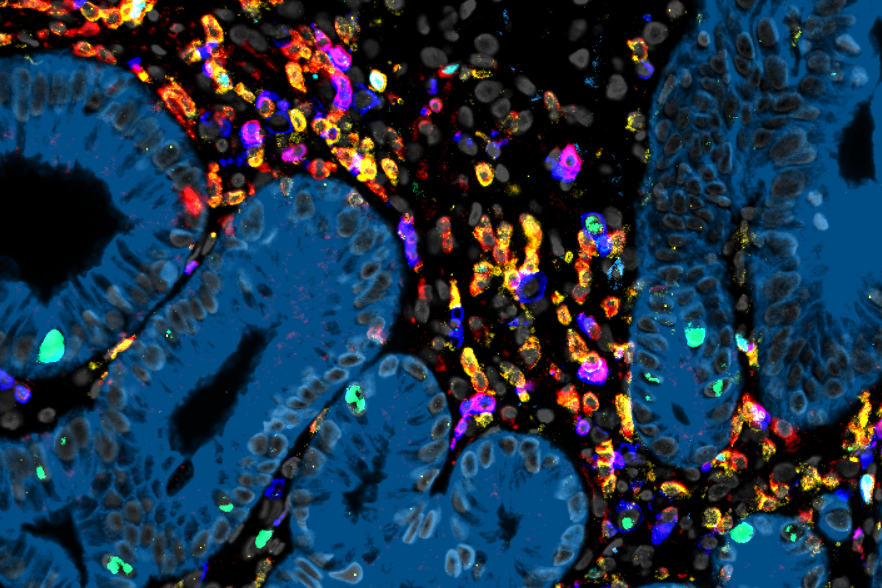
High-quality multiplexed staining of FFPE colon tissues with ChipCytometry
Adaptive T-cell responses arise from a polyclonal repertoire of naïve precursor cells. However, T-cell therapy in the context of TCR engineering usually comprises a single TCR for disease treatment. The combination of several T-cell clonotypes with different avidities and different landscapes of cross-reactivity against related targets provides flexibility in the immune response and can enhance protective capacity. Enabling polyclonality in immune responses is important in effective vaccination strategies.
Each individual harbors an enormous diversity of unique TCR sequences. Regarding protective immunity, this diversity is also necessary in order to mount effective T-cell responses against an unpredictable set of target epitopes derived from pathogens or tumor cells. TCR diversity is restricted by thymic selection processes, including the recognition of MHC and available space for maintaining T cells in an individual. However, in a similar manner epitope diversity is restricted by the capacities to bind to a specific MHC, chemical stability as well as the presence of limited key residues in an epitope that are exposed to TCR recognition. Current estimates of TCR and epitope diversity predict a high likelihood to mount an effective immune response against any given target, or in other words: there is a high probability of finding a high avidity clonotype in a naïve repertoire for any specificity.
New technologies enabled access to large-scale TCR sequencing data and will improve further in the future. However, screening TCR functionality of hundreds of thousands of TCRs in a high-throughput manner is still not feasible. We aim to understand how we can efficiently predict TCR specificity from the TCR sequence alone and combine this with the identification of high avidity clonotypes. TCR sequence data from naïve T-cell repertoires could serve as an invaluable source for rapid identification of high avidity TCRs for cellular therapy. In order to achieve this goal, we are generating large-scale TCR libraries of experimentally validated epitope specificities to elucidate TCR recognition in different contexts.
Personnel

Related publications
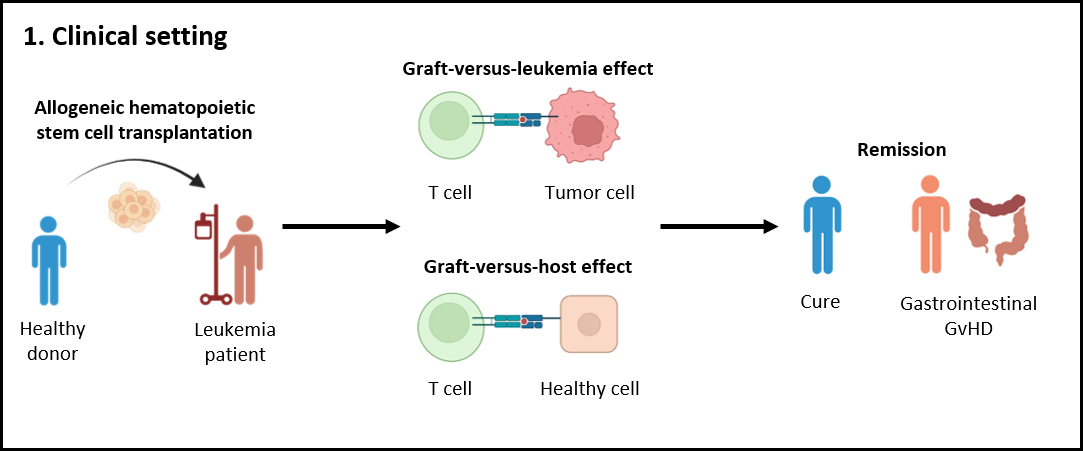
Allogeneic hematopoietic stem cell transplantation (aHSCT) holds promise as a treatment option for hematological malignancies. This procedure involves the removal of a patient's immune system through myeloablative conditioning to eliminate tumor cells, followed by the infusion of healthy donor cells. One of the significant advantages of aHSCT is the graft-versus-leukemia (GvL) effect, wherein the donor cells mount an immune response against any remaining or recurring cancer cells in the patient. However, this approach may lead to complications like graft-versus-host disease (GvHD), where the donor cells also target the recipient's healthy cells. While GvHD has been associated with a reduced risk of leukemic relapses, it can be a life-threatening complication, underscoring the critical need to steer the immune response towards GvL while preventing GvHD.
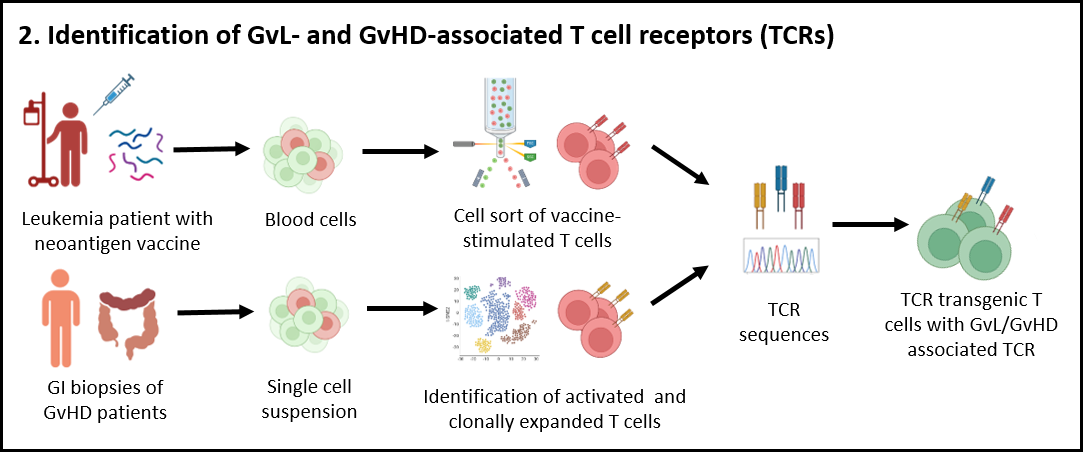
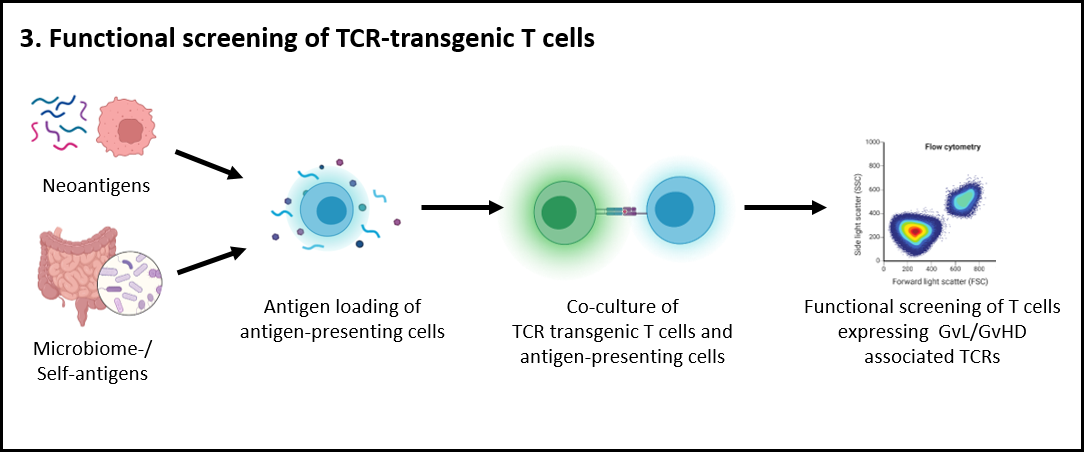
In pursuit of enhancing leukemia-specific donor cells, novel patient-personalized cancer vaccines have emerged. This approach involves identifying tumor-specific genomic mutations that can be translated into proteins and presented on the patient's tumor cells as neoantigens. These neoantigens are then synthesized and administered as a vaccine. Collaborators in Tübingen have successfully applied this method in treating young leukemia B-cell cancer patients, and observed the emergence of putative vaccine-specific T cells. Our current objective is to identify the T-cell receptors (TCRs) expressed by these cells and analyze their specificity and functionality. This investigation will validate the vaccine's ability to induce a neoantigen-specific T cell response and evaluate whether the quality of this response can strengthen the GvL effect.
Furthermore, recent studies have established a connection between GvHD and the patients' microbiome. Patients with higher microbiota diversity show reduced susceptibility to GvHD, and fecal microbiota transplantation (FMT) has demonstrated success in curing steroid-resistant GvHD. Building upon our prior research utilizing single-cell RNA sequencing and in situ multiplexed tissue imaging via ChipCytometry of gastrointestinal biopsies, we have discovered that severe GvHD patients exhibit lower suppressive capacity of regulatory T cells and a higher fraction of clonal expansion of CD8+ T cells compared to the control group without GvHD. We now aim to investigate the antigens recognized by these putative GvHD-driving T cells and explore the immunomodulatory effects of FMT and conventional GvHD treatment options.
Personnel

Related publications
Jarosch et al., Cell Rep Med, 2023
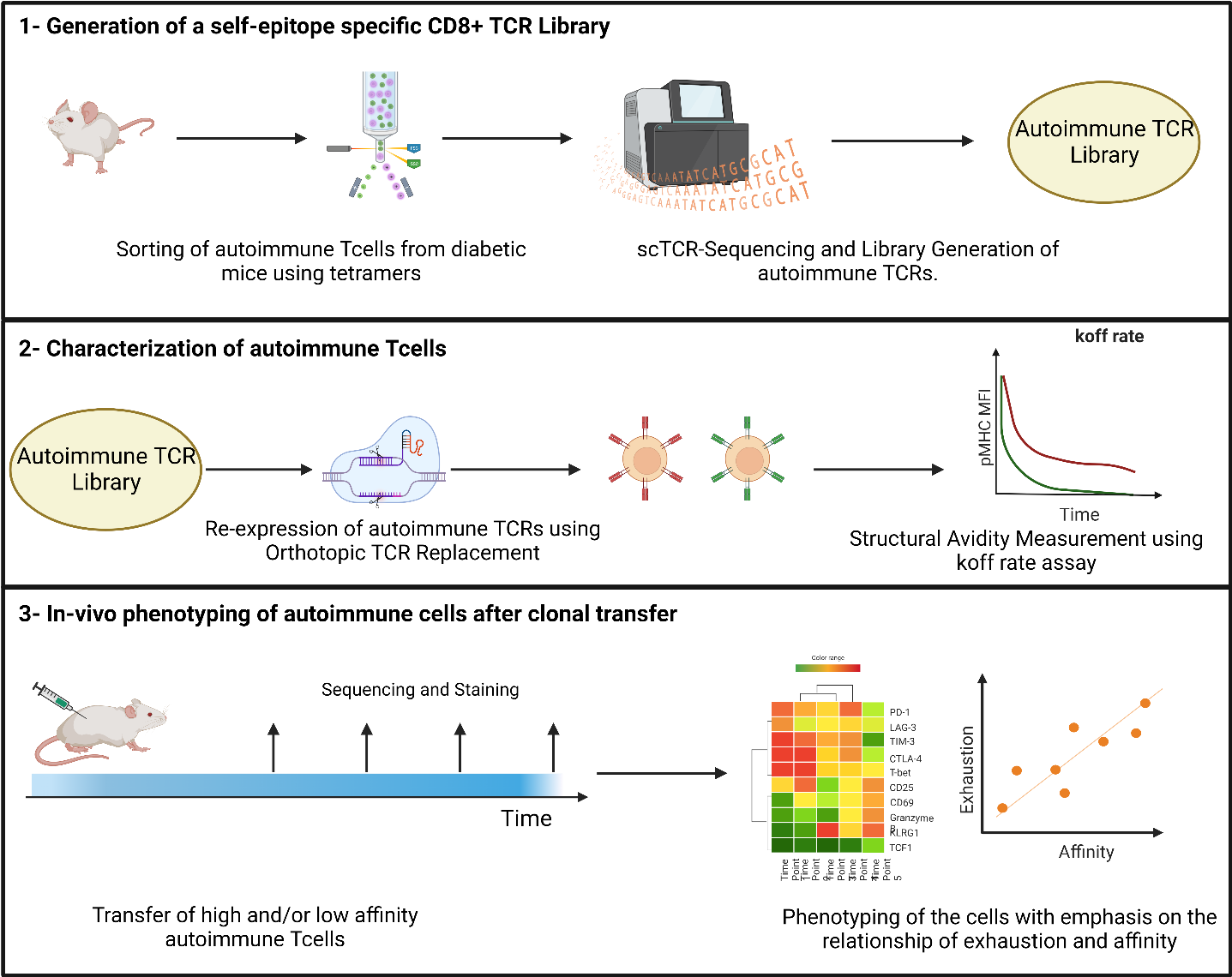
While CD8+ T-cells are the cornerstone of fighting off infections and tumors, they are also the drivers of multiple autoimmune disorders, such as type-I diabetes. CD8+ T-cells infiltrate the pancreas and destroy the insulin-secreting cells over months to years, leading to insulin-deficiency and hyperglycemia with severe vascular consequences. Despite chronic antigenic exposure, autoimmune CD8+ T-cells appear to remain more functional and cytotoxic than CD8+ T-cells in the context of tumors or chronic infections. One explanation for this divergence could be a differential TCR repertoire comprising different avidities between autoreactive cells and T cells targeting tumor or pathogen related antigens. Using single-cell RNA and TCR-sequencing, we want to sequence the TCR repertoire against the model epitope SIINFEKL, derived from Ovalbumin, in an autoimmune versus an infectious setting. The sequenced TCRs will be re-expressed in reporter cell lines and further characterized for structural and functional avidity. This way we aim to gain more insight in how the autoimmune response is initiated and maintained, paving the way for future therapies to stop the chronicity of the autoimmune destruction.
Personnel


Related publications
The emergence of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic and the resulting Coronavirus disease 2019 (COVID-19) has massively impacted everyday life. While some risk factors like cardiovascular or pulmonary comorbidity, obesity and male gender are correlated with a higher risk of severe COVID-19 disease, in many cases the cause of severe disease cannot be determined. Even though many efforts have been undertaken to get a better understanding of the pathophysiology of the breadth of disease penetrance, up until now, an extensive explanation for severe COVID-19 disease in previously healthy patients is still missing.
In previous work, we identified a pre-existing infection with the human cytomegalovirus (CMV) as risk factor for developing severe COVID-19 disease, especially in non-geriatric patients. In addition, we could show that CMV seropositivity is associated with a higher prevalence of comorbidities that are known risk factors for severe COVID-19 disease. CMV is a herpesvirus that causes a latently persisting infection that is known to reshape the immune repertoire by creating an inflationary memory T cell response. How the memory CMV-specific T-cell repertoire shapes the immune response against SARS-CoV-2 is subject of our current investigations. A potential mechanism of this might be the cross-reactivity of CMV-primed T-cell responses towards SARS-CoV-2 epitopes. Understanding the underlying causes of severe COVID-19 disease, especially in individuals previously considered healthy, is of high relevance for reducing morbidity and mortality and improving treatment strategies.
Personnel